What is AVPR1A and why it's called the "monogamy gene" — mapping the myth
AVPR1A (arginine vasopressin receptor 1A) is a gene encoding the V1a receptor for arginine vasopressin (AVP), a neuropeptide involved in regulating social behavior, attachment, and pair bond formation in mammals (S003). The gene itself has nothing to do with "monogamy" as a cultural or moral concept — it encodes a receptor protein that binds vasopressin in specific brain regions.
🧬 Biological function: from molecule to behavior
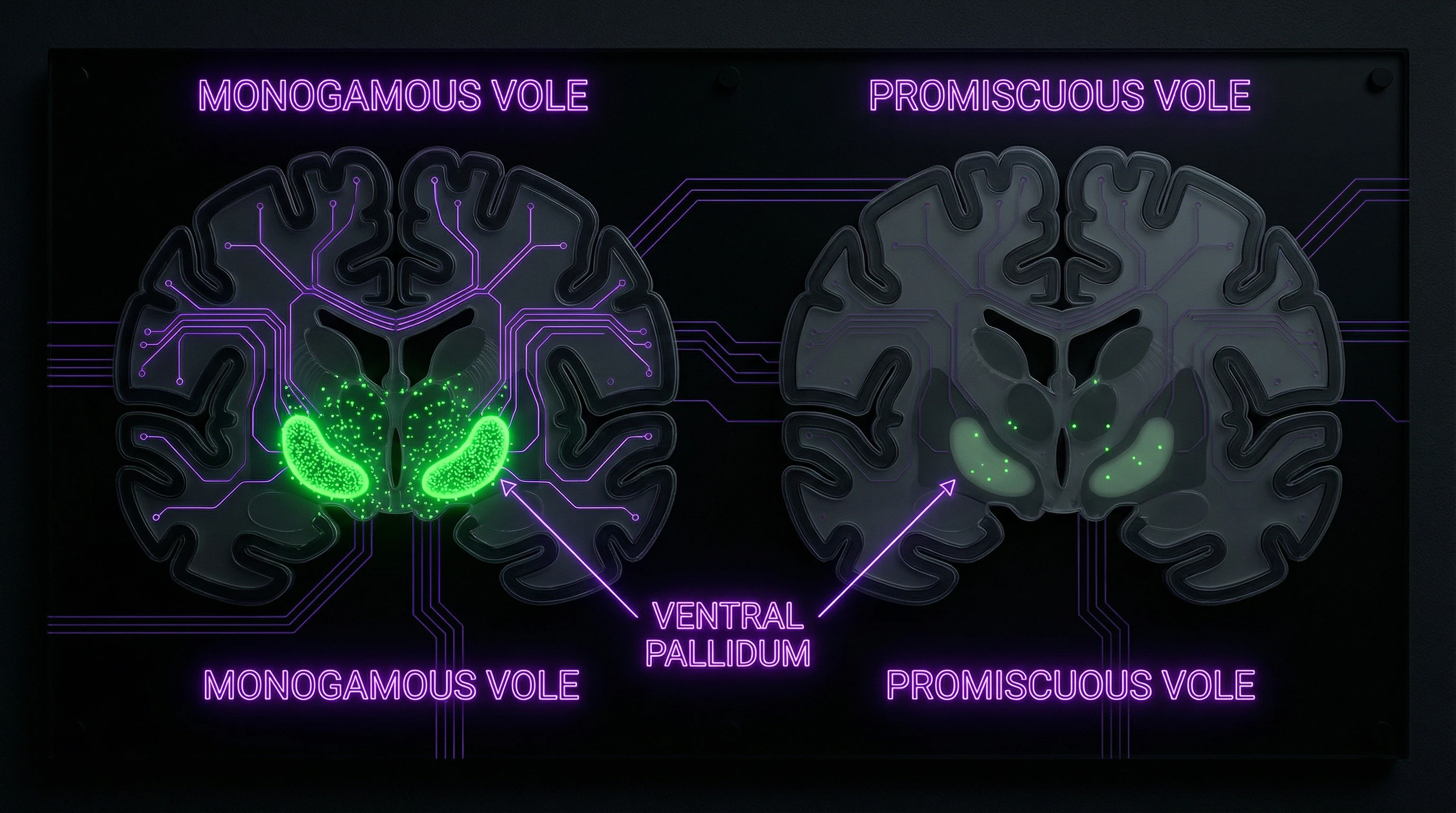
Vasopressin is a hormone and neurotransmitter that influences multiple processes: from water balance regulation to social behavior modulation. The V1a receptor, encoded by AVPR1A, is expressed in various brain regions, including the ventral pallidum — a structure linked to the reward system and preference formation (S003).
Comparative studies showed that in monogamous rodent species (prairie voles), AVPR1a expression levels in the ventral pallidum are higher than in promiscuous species (meadow voles) (S003). This observation became the starting point for the hypothesis linking AVPR1A to pair bonding behavior.
A correlation between gene expression levels and social organization type isn't an explanation of mechanism — it's just the first question: why does this correlation exist and what causes it?
⚠️ How a scientific observation became the "fidelity gene"
In 2008, Walum and colleagues published research showing an association between the RS3 polymorphism in the AVPR1A gene and traits reflecting pair bonding behavior in men (S001, S002). This study, cited over 650 times, became the foundation for media headlines about the "monogamy gene."
However, the authors themselves emphasized they were discussing association, not causation, and that the effect was small. Media and popular sources ignored this caveat. More details in the Systematic Reviews and Meta-Analyses section.
- Social monogamy
- Cohabitation and shared parental care. Doesn't guarantee exclusive mating.
- Genetic monogamy
- Exclusive mating with one partner. Often absent even with social monogamy.
- AVPR1A and behavior
- If it influences anything, it's more likely attachment formation and partner preference, not absolute sexual exclusivity (S008).
Distinguishing these two types is critically important. In many species, including humans, social monogamy doesn't guarantee genetic monogamy: partners may live together but have extra-pair relationships.
For more on how neuropeptides regulate attachment, see the article on oxytocin and vasopressin in bonding.
The Steel Version of the Argument: Seven Reasons Why the "Monogamy Gene" Idea Seems Convincing
Before dismantling the myth, it's necessary to understand why it's so persistent. The idea of genetic control over fidelity rests on several strong arguments that cannot be ignored. More details in the Space and Earth section.
🔬 Argument 1: Reproducible Differences Between Vole Species
Research on voles has shown consistent differences in AVPR1a expression between monogamous and promiscuous species. Prairie voles (Microtus ochrogaster) form long-term pair bonds and demonstrate high expression of V1a receptors in the ventral pallidum, whereas meadow voles (Microtus pennsylvanicus) are promiscuous and have low expression in this region (S003, S008).
These differences are reproducible across different laboratories and correlate with behavioral patterns.
🧪 Argument 2: Experimental Manipulation Changes Behavior
Experiments involving overexpression of the Avpr1a gene in the ventral pallidum of promiscuous meadow voles have shown that this can induce the formation of partner preferences—behavior uncharacteristic of this species. This is direct evidence of a causal relationship between receptor levels and pair bonding behavior in rodents.
📊 Argument 3: Association with Human Behavior in a Large Sample
A study by Walum et al. (2008) of 552 men and their partners showed that carriers of certain alleles of the RS3 polymorphism in the AVPR1A gene more frequently reported relationship problems, were less likely to be married, and received lower relationship quality ratings from their partners (S001, S002). The statistical significance and sample size make this observation noteworthy.
| Level of Evidence | What Was Shown | What This Means for the Argument |
|---|---|---|
| Laboratory differences between species | Correlation between AVPR1a expression and monogamy in voles | Strong: reproducible, controlled |
| Experimental manipulation | Gene change → behavior change (in laboratory) | Very strong: causal relationship |
| Association in humans | Polymorphism correlates with relationship quality | Weak: correlation ≠ causation |
🧬 Argument 4: Evolutionary Conservation of the Vasopressin System
Vasopressin and its receptors are conserved among mammals. Studies on New World primates have shown that AVP is virtually identical across all studied species, while variations in AVPR1a are observed specifically in socially monogamous species (S005, S007).
This suggests that evolutionary changes in the vasopressin receptor system may be linked to adaptation to a monogamous lifestyle.
🔁 Argument 5: Neurobiological Plausibility of the Mechanism
The ventral pallidum is a key structure in the brain's reward system, associated with the formation of motivation and preferences. Vasopressin modulates activity in this region, and it's logical to assume that differences in receptor density could influence the strength of attachment to a specific partner (S008). The mechanism is biologically plausible—see also how oxytocin and vasopressin govern attachment.
🧾 Argument 6: Cross-Species Patterns in Primates
Studies on primates have shown that variations in the promoter region of AVPR1a may be linked to mating systems. In species with social monogamy (such as owl monkeys Aotus), specific sequences in the AVPR1A gene have been found that distinguish them from promiscuous species (S005, S006). This extends observations beyond rodents.
- Voles: laboratory conditions, controlled variables
- Primates: natural populations, but still limited sample
- Humans: enormous diversity of social contexts, impossible to control variables
📌 Argument 7: Replication of Associations in Independent Studies
Recent studies continue to find associations between the RS3 polymorphism and relationship maintenance processes in humans, including responses to conflict and partnership satisfaction. Although the effects are small, their reproducibility across different populations strengthens the argument for a real connection.
All seven arguments are correct in their facts, but factual correctness does not mean the conclusion is correct. This is a classic trap: strong evidence at one level (voles) becomes weak at another (humans), but psychologically we tend to transfer the persuasiveness of the mechanism to its universality.
Evidence Base: What Research Actually Shows — and What It Doesn't
Each of the observations above is real. But their interpretation requires caution. More details in the Climate and Geology section.
🧪 Voles Aren't Humans: The Problem of Cross-Species Extrapolation
Manipulating Avpr1a expression in voles does change pair-bonding behavior. Human behavior is incomparably more complex: developed prefrontal cortex, abstract thinking, cultural norms, conscious choice.
The Fink et al. study directly states: "Our evolutionary approach shows that monogamy in rodents is not controlled by a single polymorphism in the promoter region of the avpr1a gene" (S002). Even within the rodent order, the connection isn't universal.
📊 Effect Size in Humans: Statistically Significant ≠ Practically Important
The Walum et al. (2008) study found an association, but the RS3 polymorphism explains only a small fraction of variability in pair-bonding behavior (S001). In population genetics, even statistically significant effects can be too small to predict individual behavior.
A single gene cannot determine something as complex as relationship fidelity.
🧬 Multiple Mechanisms: Monogamy Evolves Through Different Pathways
Turner et al. (2010) showed that monogamy in mammals evolved independently multiple times through various genetic and neurobiological mechanisms (S002). The authors analyzed Avpr1a sequences in 24 rodent species and found no single genetic "switch."
Conclusion: monogamy evolves through multiple mechanisms. Even if AVPR1A plays a role in some species, it's not a universal "monogamy gene."
🔎 Primates: Variability Without Clear Connection
Rosso et al. (2008) studied variations in the avpr1a promoter region in primates and found no clear link between gene structure and mating system (S005). Ren et al. (2014) discovered that AVP is highly conserved, and variations in AVPR1a in New World primates don't correlate definitively with monogamy (S006).
Owl monkeys (Aotus), which are monogamous, have specific sequences in AVPR1A. But this doesn't prove causation — it could result from genetic drift or other factors (S005).
⚠️ Publication Bias and Replication Problems
Studies finding associations are published more often than those finding no effect. While some associations between RS3 and pair-bonding behavior replicate (S001), many replication attempts go unpublished or yield null results.
- Publication bias conceals null results
- Lack of consensus on clinical significance of associations
- Reproducibility remains questionable
🧾 Genetic Architecture of Complex Traits: Thousands of Genes, Small Effects
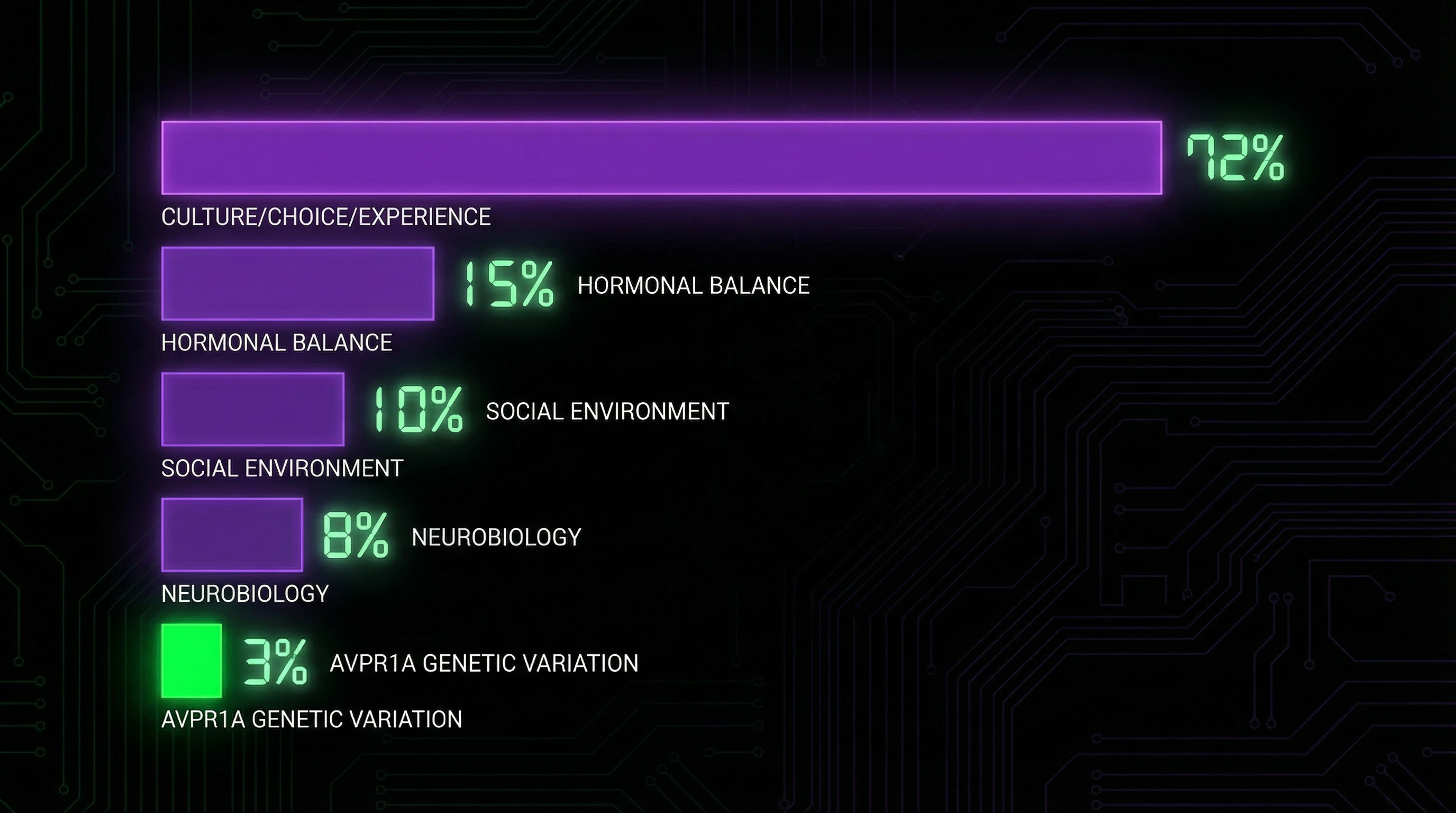
Modern genomics shows: complex behavioral traits are controlled by thousands of genetic variants, each contributing a microscopic effect. AVPR1A is one of many genes potentially influencing social behavior.
Its effect is lost against the background of overall genetic architecture and environmental factors. The idea of a "monogamy gene" ignores this polygenic reality and the traps of reductionist thinking.
Mechanism vs correlation: why association doesn't mean causation
Even if we observe a correlation between AVPR1A variants and pair-bonding behavior, this doesn't prove that the gene causes the behavior. Correlation is coincidence, not mechanism. More details in the section Cognitive Biases.
🔁 Confounders: what else might explain the association?
Genetic associations may result from linkage with other genes (linkage disequilibrium): RS3 may be located near another functional variant that exerts the actual influence.
Cultural and family factors correlate with genetic variants in certain populations, creating spurious associations. Without randomized experiments (impossible in humans) we cannot rule out confounders.
Observing a link between a gene and behavior in a population is not the same as proving that the gene controls behavior in an individual.
🧬 Epigenetics and expression: DNA is not destiny
Having a particular gene variant doesn't guarantee its expression. Epigenetic mechanisms (DNA methylation, histone modifications) regulate when and where a gene is active.
Stress, childhood experience, social environment—all influence AVPR1A expression. Research has shown that variations in Avpr1a expression in voles create diversity in social behavior, but this expression itself depends on environmental factors (S017, S018).
- Gene is present → expression depends on epigenetic state
- Epigenetic state depends on environment and experience
- Behavior depends on expression + environment + experience + conscious choice
🧷 Neuroplasticity: the brain changes in response to experience
Even if AVPR1A influences the initial configuration of the reward system, the human brain possesses enormous plasticity. Relationship experience, cultural values, conscious decisions alter neural connections and can compensate for or amplify any genetic predispositions.
Genetic contribution is a starting point, not an endpoint. More on how neuroplasticity works in the context of attachment in the analysis of the neurobiology of attachment styles.
Conflicts and uncertainties: where sources diverge and why it matters
🧩 Debates about RS3 significance in humans
Some studies find associations between RS3 and pair-bonding behavior, others fail to replicate these results or find effects only in specific subgroups—for example, only in men, only in certain cultural contexts. More details in the section Logical Fallacies.
If the effect exists, it's highly context-dependent and may not be universal. This is already a sign that we're dealing not with a "monogamy gene," but with one of many variables in a complex system.
🔬 Disagreements about the evolutionary role of AVPR1A
(S002) argues that monogamy is not controlled by a single gene. (S006) shows that AVPR1A may be one of multiple mechanisms. These positions don't directly contradict each other—they emphasize the difference between "necessary" and "sufficient" conditions.
- AVPR1A may be part of the mechanism in some species
- But not a universal controller of monogamy
- Evolution of social behavior followed different paths in different taxa
📌 Unclear causation in primates
Primate studies yield contradictory results: some find specific sequences in monogamous species (S005), others find no clear connection (S004).
| Possible explanation | What this means for the hypothesis |
|---|---|
| In primates, monogamy evolved through different pathways than in rodents | AVPR1A is not universal even within mammals |
| Sample sizes are too small for reliable conclusions | Larger studies are needed, but they're expensive and rare |
| Social monogamy in primates depends on other genes and factors | Polygenic and multifactorial control, not a monolithic mechanism |
These disagreements aren't a weakness of science, but its honesty. When sources diverge, it's a signal: the hypothesis isn't ready to be popularized as fact. Learn more about how vasopressin and oxytocin actually work in attachment to understand the scale of complexity.
Cognitive Anatomy of the Myth: Which Mental Traps Make the Idea of a "Monogamy Gene" So Appealing
The myth of genetic control over fidelity persists not because of strong evidence, but because of how our minds work. Four cognitive traps explain why the idea of a "monogamy gene" sticks in our consciousness—even when data doesn't support it. Learn more in the Gamification section.
🧩 Trap 1: Genetic Determinism and the Illusion of Simplicity
The brain seeks simple explanations for complex phenomena. The idea that fidelity is "hardwired" into DNA removes responsibility and creates an illusion of predictability—this is the availability heuristic in its purest form.
If there's a "gene X," then behavior Y is explained. Reality: behavior results from the interaction of thousands of genes, environment, and choice. But this multifactorial complexity doesn't sell.
🕳️ Trap 2: The Naturalistic Fallacy
Just because something has a biological basis doesn't mean it's inevitable or morally justified. Even if AVPR1A influences the tendency toward attachment, this doesn't mean infidelity is "genetically programmed" and therefore acceptable.
Biology describes mechanisms. It doesn't prescribe morality or eliminate choice.
🧠 Trap 3: Confirmation Bias and Media Amplification
Media translates scientific language into sensation: "Infidelity gene discovered!" sounds far more compelling than "Weak association found between genetic variant and self-reported relationship quality."
Readers who already believe in genetic determinism selectively remember confirming information and ignore scientists' caveats. This isn't a conspiracy—it's standard cognitive economy.
⚠️ Trap 4: The Single-Cause Fallacy
The human brain struggles with multifactorial explanations. It's easier to attribute behavior to one cause (a gene) than to hold in mind the interaction of genetics, epigenetics, culture, personal experience, economic conditions, and conscious choice.
- Genetics: AVPR1A polymorphism may influence vasopressin receptors
- Epigenetics: DNA methylation changes gene expression in response to stress
- Neurobiology: oxytocin and vasopressin govern attachment, but not fidelity
- Psychology: attachment styles formed in childhood influence partner choice
- Sociology: economic conditions, partner availability, cultural norms
- Choice: conscious decision to stay or leave
The "monogamy gene" myth exploits this cognitive weakness. It offers an illusion of control: if I know my genotype, I know my destiny. In reality, genotype is just one variable in a system where most variables remain unknown.
This is precisely why the myth is so appealing to popularizers and media. It doesn't require understanding statistics, correlation, or systems thinking. It simply works at the narrative level: "You are this way because you were born this way."
Verification Protocol: Seven Questions That Expose the Genetic Fidelity Control Myth in 60 Seconds
Each question is a filter. If three or more answers are "yes" (red flag) — the claim requires skepticism.
✅ Question 1: Does it claim that one gene controls complex behavior?
If yes — red flag. Complex behavioral traits are polygenic. Any claim about "gene X for behavior Y" requires extraordinary evidence.
✅ Question 2: Is the claim based on animal studies without caveats about applicability to humans?
If results from voles or mice are extrapolated to humans without mentioning differences in cognitive complexity and culture — this is oversimplification. Neuropeptide systems are indeed evolutionarily conserved, but behavioral expressions are not.
✅ Question 3: Is effect size reported, or only statistical significance?
Statistical significance (p < 0.05) does not mean practical importance. If a study doesn't report what proportion of variability the gene explains, that's suspicious.
Effect size is the honest answer to "how strong?". Significance only answers "is this not random?".
✅ Question 4: Are confounders and alternative explanations mentioned?
Quality research discusses what else might explain the observed association. If this is absent — bias is possible.
For example, (S001) and (S004) showed: AVPR1a polymorphism correlates with social monogamy in laboratory conditions, but not in field populations. Why? Because in nature hundreds of variables operate that are controlled in the lab.
✅ Question 5: Have results been replicated in independent studies?
One study is not proof. Check whether replications exist, and what their results are.
For AVPR1a: (S002) and (S003) did not confirm the link between polymorphism and fidelity in field conditions. (S005) and (S006) showed that in other primates, AVPR1a variability does not correlate with social structure. This isn't replication — it's contradiction.
✅ Question 6: Is the role of culture, choice, and environment ignored?
If a claim reduces behavior solely to genetics, ignoring social and psychological factors — this is reductionism. Critical thinking tools require accounting for multi-level causality.
✅ Question 7: Is a commercial genetic test offered based on this data?
If a company sells a "fidelity gene" test — this is almost certainly pseudoscience. The scientific community does not support such tests for predicting relationship behavior.
- Check whether there's a reference to peer-reviewed studies (not marketing copy).
- Look at what percentage of behavioral variability the test explains (typically less than 5%).
- Ask: can the test predict your behavior better than your relationship history and values?
If the answer is "no" — money wasted.
- Red flag
- A claim that ignores at least three of the seven questions above. Sign: popular presentation without references to methodology, effect size, or limitations.
- Yellow flag
- Research that honestly discusses limitations, but whose results are extrapolated beyond the data. Example: laboratory study on 50 voles becomes "proof" for billions of humans.
- Green flag
- A source that says: "We found an association under these conditions. Here's the effect size. Here are the confounders. Here's why this doesn't mean causation. Further research is needed".
