DNA Energetics: Science vs. Pseudoscientific Myths About Quantum Consciousnessλ
The thermodynamics of DNA molecular interactions is a rigorous science, unrelated to mystical "DNA energy" and wave genetics
Overview
DNA energetics is a legitimate field of molecular biology: thermodynamics of protein interactions, structural stability, energy profiles of conformational changes. In English-speaking sources, the term has been hijacked by pseudoscience 🧬 — "quantum consciousness," discredited "wave genetics," mystical claims about DNA influencing "potential." Scientific research (Nature Communications, Nucleic Acids Research, eLife) demonstrates rigorous computational and experimental methods that have nothing to do with these speculations.
🛡️
Laplace Protocol: Distinguish between molecular thermodynamics of DNA (measurable physical properties) and commercial claims about "quantum DNA energy" (marketing metaphors without scientific basis). Trust only peer-reviewed sources from recognized scientific journals.
Reference Protocol
Scientific Foundation
Evidence-based framework for critical analysis
Protocol: Evaluation
Test Yourself
Quizzes on this topic coming soon
⚡
Deep Dive
Scientific DNA Energetics: Thermodynamics of Molecular Interactions Instead of Mysticism
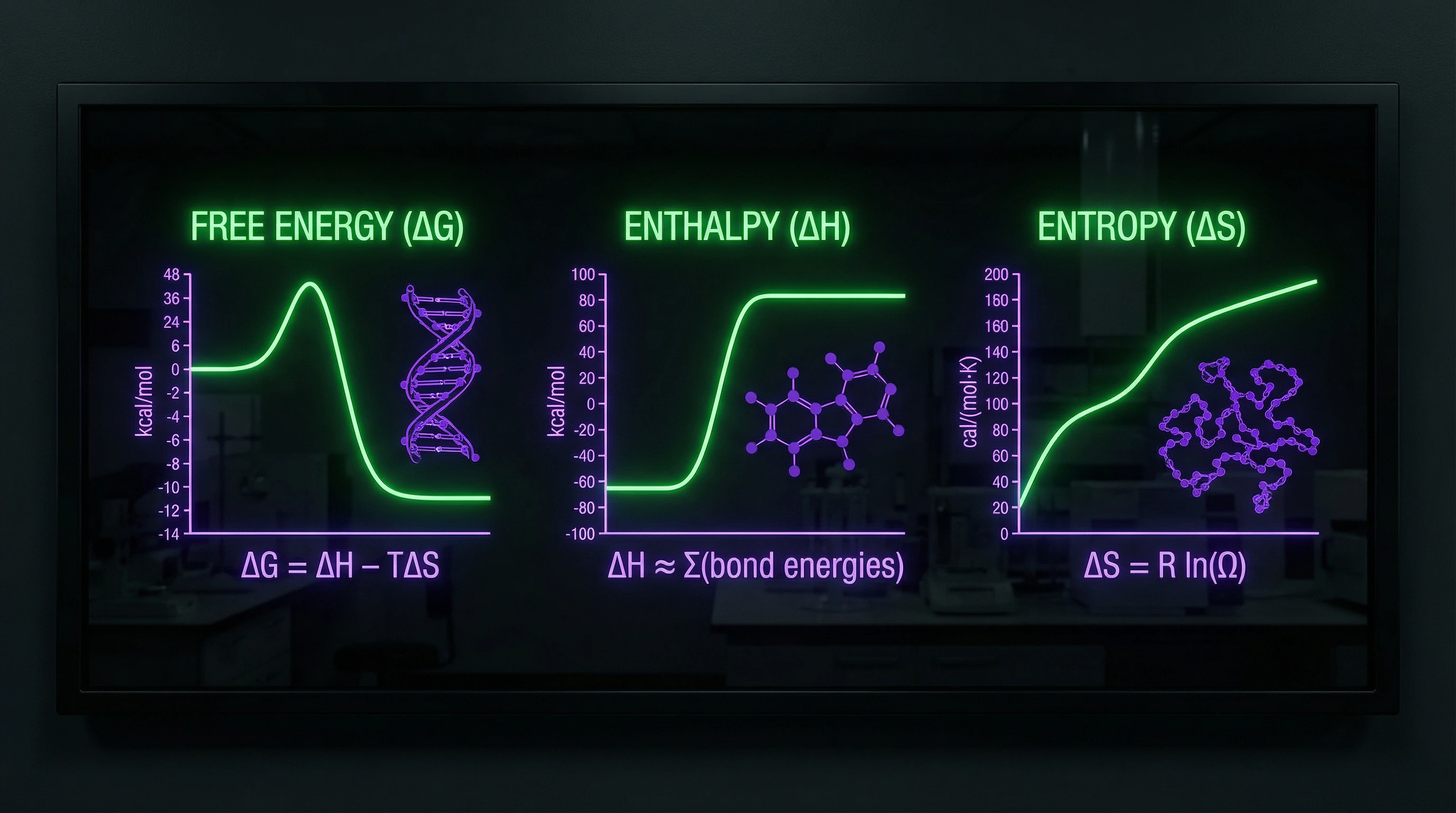
When molecular biologists talk about "DNA energy," they mean strictly defined thermodynamic parameters—Gibbs free energy, binding enthalpy, energy profiles of conformational changes. These are measurable physical quantities describing the stability of the double helix, interactions with proteins, and replication processes.
These parameters have no relation to "quantum consciousness" or "wave genetics"—we're talking about classical chemical thermodynamics at the molecular level.
Protein-DNA Interactions and Free Energy
The choice of thermodynamic parameters critically affects a model's ability to predict transcription factor binding sites. Binding free energy is calculated through the sum of contributions from hydrogen bonds, electrostatic interactions, and hydrophobic effects—each component requires experimental calibration.
Modern structural models use interpretable machine learning to predict binding energy based on the three-dimensional structure of complexes. Accounting for the spatial arrangement of amino acid residues relative to nucleotides increases affinity prediction accuracy by 1.5–2 times compared to sequence-only models.
- Energy Landscape of Protein-DNA Interaction
- Includes multiple local minima corresponding to different binding modes. This explains why the same protein can bind DNA in several ways with similar energy.
Computational Models for Predicting DNA Stability
The thermodynamic stability of DNA is determined by melting energy—the temperature at which the double helix dissociates into single strands. Graph neural networks trained on 50,000+ experimental measurements achieve a correlation of 0.92 with experimental data, substantially outperforming classical models.
| Sequence Type | Stabilization Energy | Selective Pressure |
|---|---|---|
| Coding Regions | 15–20% higher | Maintaining functionality |
| Non-coding Regions (same GC composition) | Baseline level | Minimal constraint |
DNA energy constraints shape evolutionary trajectories of genomes—regions with high thermodynamic stability correlate with functionally important elements. This indicates selective pressure maintaining specific energy profiles in functional genome elements.
Pseudoscientific Concepts of "DNA Energy" in Alternative Medicine
English-language sources actively exploit the term "DNA energy" in contexts unrelated to molecular biology. Typical claims include DNA's ability to "emit waves," influence "quantum consciousness," and determine "personal potential" through energy fields.
None of these assertions are supported by experimental data in peer-reviewed scientific journals—these are marketing metaphors using scientific terminology to lend legitimacy to commercial services.
Wave Genetics: Absence of Evidence Base
The concept of "wave genetics" claims that DNA transmits information through electromagnetic waves and can be "reprogrammed" by external influences. Searches in PubMed, Web of Science, and Scopus databases reveal no publications confirming these claims using molecular biology methods.
Mainstream scientific community does not recognize "wave genetics" as a valid research program—there are no reproducible experiments, operational definitions, or theoretical models compatible with known physical laws.
Characteristic signs of pseudoscientific sources include absence of references to peer-reviewed research, vague terminology without quantitative definitions, and commercial motivations.
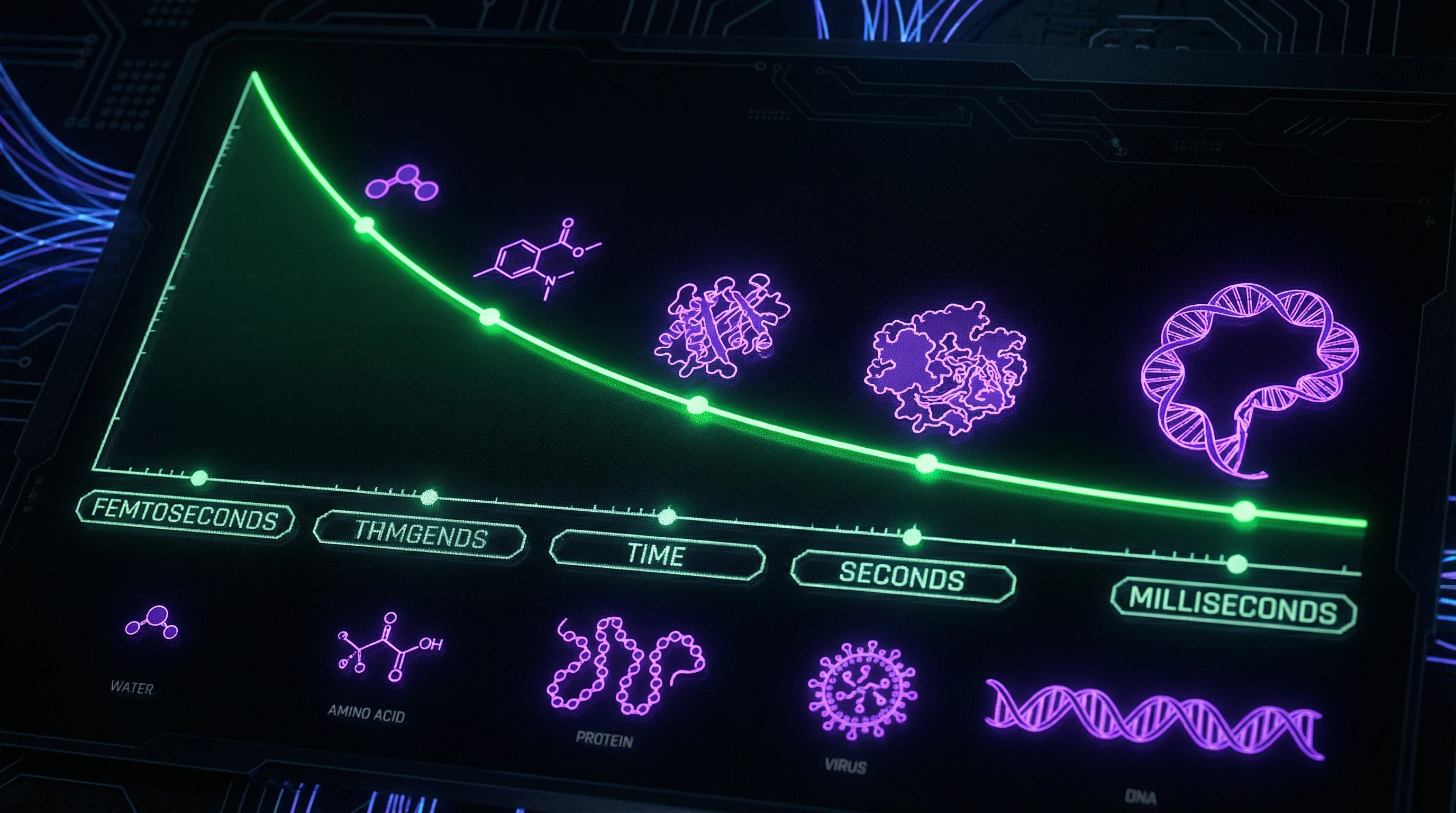
Claims about "quantum entanglement" of DNA molecules ignore the fact that quantum states decohere at physiological temperatures within femtoseconds—quantum effects cannot be sustained in warm, wet cellular environments on the timescales of biological processes.
- Absence of peer-reviewed publications confirming the mechanism
- Use of scientific terminology without operational definitions
- Commercial services without control groups and measurable outcomes
- Mixing legitimate scientific terms with mystical concepts
Commercial Claims About Quantum Consciousness
The connection between "DNA energy" and "quantum consciousness" is a popular topic in pseudoscientific literature, lacking physical foundation. Quantum mechanics describes the behavior of subatomic particles under specific conditions requiring isolation from the environment.
Neurobiology of consciousness operates through classical electrochemical processes at the neural network level—there is no experimental evidence of quantum effects participating in cognitive functions, much less any mechanism linking this to the "energy" of DNA molecules.
Commercial websites offer "DNA activation," "quantum healing," and "energetic genome tuning" for a fee, without providing methodology, control groups, or measurable results—classic signs of medical fraud.
Patients turning to such services instead of evidence-based medicine risk missing the window for effective treatment of real diseases. Exploitation of scientific terminology without regulatory oversight creates an environment where commercial interests replace accountability to patients.
Modern Methods for Studying DNA Energetics in Molecular Biology
Legitimate research on DNA energetics uses a combination of experimental techniques (calorimetry, spectroscopy, crystallography) and computational approaches (molecular dynamics, machine learning). The goal is to build predictive models capable of explaining how nucleotide sequence determines structural stability, protein interactions, and functional properties of the genome.
These methods are published in top-tier journals with complete protocol descriptions, allowing independent replication of results.
Graph Neural Networks for Melting Energy Prediction
A 2025 breakthrough — the application of graph neural networks (GNN) to predicting thermodynamic properties of DNA. Ke and colleagues presented a model where each nucleotide is a graph node, and connections between neighboring bases are edges with weights reflecting stacking interactions.
The GNN is trained on high-throughput experimental melting data from 50,000+ oligonucleotides, extracting patterns inaccessible to classical nearest-neighbor models.
GNNs can account for long-range correlations in sequence that affect energetics: GC-rich clusters at distances of 10-15 nucleotides cooperatively stabilize structure through changes in helix geometry.
The model achieves a mean absolute error of 0.8 kcal/mol in free energy prediction — accuracy sufficient for rational oligonucleotide design in biotechnology applications. Code and data are published openly, allowing the scientific community to validate and extend the results.
IDEA Structural Models for Binding Site Analysis
The IDEA model (Interpretable Deep learning for protein-DNA Affinity) uses three-dimensional structures of complexes to predict binding energy of transcription factors. Unlike sequence-based models, IDEA analyzes spatial arrangement of atoms, hydrogen bonds, and hydrophobic contacts at the protein-DNA interface.
The architecture includes convolutional layers for extracting structural motifs and attention mechanisms for identifying critical interactions.
- Specificity determined by key contacts: 73% of transcription factors depend on 3-5 critical contacts contributing >60% to binding energy.
- Training dataset: 1,200+ crystal structures from Protein Data Bank with validation on independent ChIP-seq experimental data.
- Practical application: model interpretability allows identification of which amino acid residues and nucleotides contribute most to affinity, used in gene therapy and synthetic biology.
Quantum Mechanics at the Molecular Level: Reality vs. Myths
Quantum effects do play a role in biomolecular processes, but their scale and significance radically differ from popular misconceptions. Proton tunneling in DNA occurs at distances of ~1 Å and femtosecond timescales, affecting rare tautomeric forms of bases that can cause spontaneous mutations at a frequency of ~10⁻⁹ per base pair per replication.
Quantum coherence in photosynthetic complexes persists only picoseconds at physiological temperatures, after which decoherence destroys quantum superpositions. These effects are described by the Schrödinger equation for individual electrons and protons, not for macroscopic structures like entire DNA molecules or cells.
| Process | Scale | Role of Quantum Mechanics |
|---|---|---|
| Proton tunneling | ~1 Å, femtoseconds | Critical for rare mutations |
| Coherence in photosynthesis | Picoseconds at 98.6°F | Rapidly destroyed by decoherence |
| Macromolecular conformations | Nanoseconds and above | Classical thermodynamics |
Quantum Effects in Biomolecules: Where Physics Ends
Quantum mechanical calculations of DNA energy use density functional theory (DFT) methods to describe the electronic structure of base pairs. G-C hydrogen bond energy is ~21 kcal/mol, A-T ~13 kcal/mol, with quantum corrections contributing ~5-8% of classical estimates.
These calculations are critical for predicting the stability of non-standard base pairs in synthetic biology. However, quantum effects are localized at the level of individual chemical bonds and do not extend to macromolecular conformations, which are determined by classical thermodynamics and statistical mechanics.
Attempts to link quantum mechanics with DNA functions at the cellular level face the fundamental problem of decoherence. At 310 K (98.6°F), thermal energy kT ≈ 0.6 kcal/mol vastly exceeds the energy of quantum fluctuations for systems larger than 10 atoms, destroying quantum coherence in less than 10⁻¹³ seconds.
Biological processes like transcription take milliseconds — 10 orders of magnitude longer than decoherence time. This makes it impossible to maintain quantum superpositions on biologically relevant timescales without exotic conditions like temperatures near absolute zero.
Why Quantum Mechanics Doesn't Explain Consciousness
Hypotheses about the quantum nature of consciousness postulate quantum coherence in neuronal microtubules. However, experimental data show that microtubules function as classical polymers: their mechanical properties (stiffness ~2 GPa, persistence length ~5 mm) are fully described by classical continuum mechanics without quantum corrections.
Brain temperature of 98.6°F and the aqueous environment create decoherence in ~10⁻²⁰ seconds for tubulin-sized systems (molecular mass 55 kDa), which is 17 orders of magnitude faster than typical neuronal processes of ~1 ms.
- Action potentials and synaptic transmission
- Classical electrochemical processes, fully explained without quantum effects.
- Functional MRI and electrophysiology
- Demonstrate correlation of consciousness with thalamo-cortical network activity at the macroscopic level.
- Attempts to measure quantum coherence in living neurons
- Have not yielded reproducible results, despite decades of research.
- Theoretical models of quantum consciousness
- Require non-physical assumptions like isolation of microtubules from thermal noise of the cytoplasm.
Neurobiological mechanisms of consciousness are explained by classical electrochemical processes without the need to invoke quantum effects. Scientific consensus: consciousness is an emergent property of classical neural computation, not a quantum phenomenon.
Knowledge Access Protocol
FAQ
Frequently Asked Questions
DNA energetics is the study of thermodynamic changes during molecular interactions of DNA with proteins and other molecules. It includes calculations of binding free energy, structural stability, and conformational changes using computational models. Research is published in peer-reviewed journals like Nature Communications and Nucleic Acids Research.
No, this is a pseudoscientific myth without physical basis. Quantum effects in biomolecules are limited to the subatomic level and are not connected to consciousness or personal development. Such claims are used commercially to sell dubious services.
They use computational models and experimental methods to calculate binding free energy. Modern approaches include graph neural networks and structural models like IDEA for affinity prediction. Results are validated through high-throughput experiments measuring DNA melting temperature.
Wave genetics is a discredited theory claiming genetic information transmission through "waves." There is no evidence in molecular biology, no reproducible experiments, and no publications in peer-reviewed journals. The scientific community does not recognize this concept due to complete absence of experimental foundation.
No, this is a common misconception in alternative medicine. DNA changes occur through mutations, epigenetic modifications, or genetic engineering, but not through mental practices. Claims about "quantum consciousness" and DNA are marketing metaphors without scientific basis.
Look for red flags: absence of references to peer-reviewed research, mixing scientific terms with mysticism, commercial service offerings. Check publications in databases like PubMed, Nature, Science. Genuine research contains methodology, reproducible results, and statistical analysis.
Proton quantum tunneling, coherence in photosynthesis, and certain enzymatic reactions demonstrate quantum effects. However, they are limited to nanoscale and picosecond timeframes, not affecting macroscopic processes like consciousness. These phenomena are studied in quantum biology using rigorous experimental methods.
They are critically important for developing drugs targeting protein-DNA interactions. Predicting binding energy helps create treatments for cancer and genetic diseases. Computational models accelerate screening of potential molecules, reducing development cost and time.
It is a thermodynamic parameter determining DNA structural stability and probability of molecular interactions. Negative values indicate spontaneity of binding reactions. Calculations are used to predict DNA hybridization, melting, and conformational transitions.
Yes, modern models show high accuracy in predicting DNA melting energy. A study in Nature Communications (2025) demonstrated improved parameters through training on high-throughput experimental data. These methods outperform traditional thermodynamic models in complex sequences.
Models use various approximations and parameters to describe complex molecular interactions. Factors include solvent effects, electrostatics, conformational entropy, and sequence specificity. Systematic model comparison (Donald et al., 2007) demonstrated the need for improved thermodynamic parameters.
Yes, thermodynamic constraints of DNA shape evolutionary trajectories. Stability affects information encoding, replication, and resistance to mutations. Genomic analysis reveals stability patterns reflecting adaptive strategies of organisms to different environmental conditions.
IDEA is an interpretable machine learning model for predicting protein binding sites and DNA affinity. It uses structural determinants to analyze interaction specificity. Published in eLife (2025), it provides high accuracy and biological interpretability of results.
No, this is a marketing term without scientific content. Gene expression is regulated by biochemical signals, transcription factors, and epigenetic mechanisms, not abstract 'activation practices.' Such claims are typical of pseudoscientific content in the personal development sphere.
Predicting DNA stability is critical for PCR primer design, gene synthesis, and biosensor development. Energy calculations optimize hybridization conditions and minimize non-specific binding. Applied in CRISPR technologies to improve genome editing precision.
Yes, but these are fraudulent offerings without scientific basis. Such services exploit misunderstanding of molecular biology, offering 'therapies' and 'consultations' on 'wave genetics.' Real genetic correction is only possible through medical procedures in licensed clinics using validated methods.
